Data; 2023/05/25(Thu)
Recently I have used scanpy for analyzing single cell RNA sequence (scRNA-seq). I feel staistical analysis on scanpy is less useful than Seurat, but scanpy has really nice compatibility to other cool software, such as scVelo, cellphoneDB, celltypist, and STREAM. This is why I like scanpy.
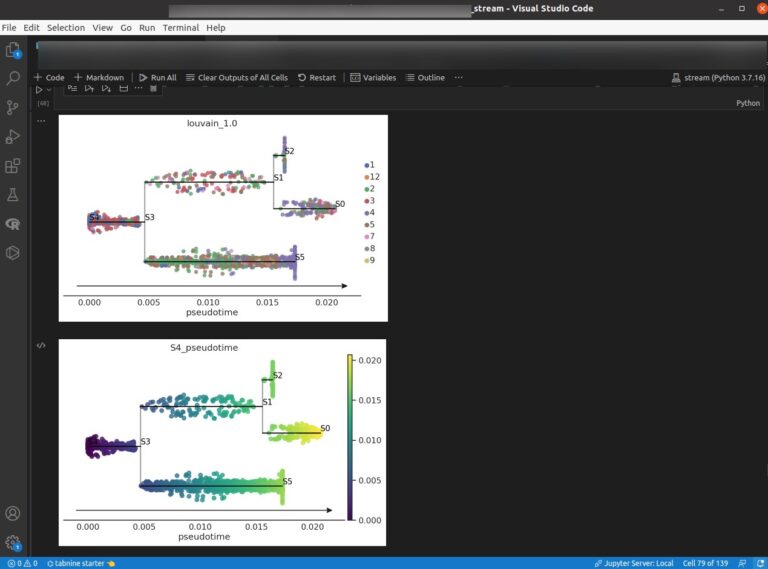
Of these softwares, STREAM performs trajectory inference and pseudotime analysis. While there are some good software for trajectory inference, such as Monocle3, Slingshot, and TradeSeq, this STREAM is more nicer than these softwares. Because pseudotime along lineage of STREAM, which is named “streamplot”, is very easy to look their path.
However, install of STREAM was super difficult for me since the install is totally stucked for some reason even though I followed instruction. However, I finally solved the problems to install STREAM. It is quite difference from the instruction, and therefore I would record my way to install STREAM.
Followed the instruction, but it did not work
At first, of course, I followed the insruction as the below. Set the repository for anaconda at first, create virtual environment for STREAM, and then install STREAM into the environment.
# this was OK. It is quite usual.
conda config --add channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge
# But, this was NOT worked.
conda create -n stream python=3.7 stream=1.0 jupyterHowever, the code describing above was totally not working. I mean, searching the packages took time forever. I think it stucked. After that, I tryed to use the code the below.
# does NOT work!!
conda install stream
# Too slow, and it does NOT work!!
conda install notebook ipykernel
# does NOT work!!
conda install stream==1.1
# does NOT work!!
conda install stream==1.0
# does NOT work!!
conda install -c bioconda stream==1.1
# does NOT work!!
conda install -c bioconda stream==1.0
# does NOT work!!
pip install stream==1.1I really stucked. While I started to crawling on Google, I could not find any useful information. but I tried another way to install STREAM.
Try not to use conda for installing STREAM
The installation using conda is always stucked at conda-forge. Actually, the conda-forge was not required for installing of STREAM (Of course, conda-forge may be required to solve dependency, but they were not working for installing STREAM, meaning conda-forge is useless for STREAM), becuase STREAM was stored at bioconda, both of version 1.0 and 1.1. You can check which repository is stored STREAM. You can see it using the below.
conda search streamBetter to use setup.py to install STREAM
At first, I set repository for conda like the below.
conda config --add channels defaults
conda config --add channels bioconda
conda config --append channels conda-forge
conda config --set channel_priority strictThen, I installed necessary environment for analysis, like the below.
conda create -n stream python=3.7conda activate stream
conda install jupyter
conda install notebook
conda install ipykernelWhen I move to VScode and tested if install is OK, they showed some errors. So I removed these errors using the blow, and then jupyter notebook was worked on stream environment which was prepared the above.
conda install -n stream ipykernel --update-deps --force-reinstallNext, I cloned the git of developer’s page.
sudo git clone 'https://github.com/pinellolab/STREAM'I think STREAM directory will be made on your current directory. Move to STREAM directory, and run setup.py.
cd STREAM
python3 setup.py --helpAs the setup.py was run, several errors were out. These errors indicated several packages are missing. So, install these missing packages. In my case, run the below.
pip install anndata
pip install netowrkx==2.1 # see https://github.com/pinellolab/STREAM/issues/18
pip install statsmodels
pip install seaborn
pip install plotly
pip install scikit-learn
pip install umap
pip install slugify
pip install rpy2
pip install shapelyThen, run the blow.
cd STREAM
python3 setup.py installFinally, STREAM is isntalled.
This analysis is scRNA-seq, and therefore run the blow.
pip install scanpy==1.9.0Critical packages for STREAM installation
What the important for installing STREAM is version of some softwares. Following are very was very critical software to process the data with STREAM
- networkx==2.1
- scanpy==1.9.0
- matplotlib==3.2.2
networkx, version 2.1 is required for running st.plot_dimension_reduction(). Scanpy, version 1.9.0 is the most latest version that can be compatible to networkx==2.1. Do not mind if you see some warnings at installation of scanpy==1.9.0. If you want to clear these warnings at installation, probably the version of scanpy must be going down more. However, My environment is normally working with scanpy==1.9.0 with networkx==2.1. matplotlib==3.2.2 is required for st.plot_de_markers().
Packages in environment of STREAM
Following is my environment. I recorded it for my reminder.
Package Version
--------------------- -----------
anndata 0.8.0
backcall 0.2.0
backports.zoneinfo 0.2.1
certifi 2022.12.7
cffi 1.15.1
cycler 0.11.0
debugpy 1.5.1
decorator 5.1.1
diopy 0.5.5
entrypoints 0.4
fa2 0.3.5
fonttools 4.38.0
h5py 3.8.0
importlib-metadata 6.6.0
ipykernel 6.15.2
ipython 7.31.1
jedi 0.18.1
Jinja2 3.1.2
joblib 1.2.0
jupyter_client 7.4.9
jupyter_core 4.11.2
kiwisolver 1.4.4
llvmlite 0.39.1
MarkupSafe 2.1.2
matplotlib 3.2.2
matplotlib-inline 0.1.6
natsort 8.3.1
nest-asyncio 1.5.6
networkx 2.1
numba 0.56.4
numpy 1.21.6
packaging 23.1
pandas 1.3.5
parso 0.8.3
patsy 0.5.3
pexpect 4.8.0
pickleshare 0.7.5
Pillow 9.5.0
pip 22.3.1
plotly 5.14.1
prompt-toolkit 3.0.36
psutil 5.9.0
ptyprocess 0.7.0
pycparser 2.21
Pygments 2.11.2
pynndescent 0.5.10
pyparsing 3.0.9
python-dateutil 2.8.2
pytz 2023.3
pytz-deprecation-shim 0.1.0.post0
pyzmq 23.2.0
rpy2 3.5.12
scanpy 1.9.0
scikit-learn 1.0.2
scipy 1.7.3
seaborn 0.12.2
session-info 1.0.0
setuptools 65.6.3
shapely 2.0.1
six 1.16.0
slugify 0.0.1
statsmodels 0.13.5
stdlib-list 0.8.0
stream 1.1
tenacity 8.2.2
threadpoolctl 3.1.0
tornado 6.2
tqdm 4.65.0
traitlets 5.7.1
typing_extensions 4.6.0
tzdata 2023.3
tzlocal 4.3
umap 0.1.1
umap-learn 0.5.3
wcwidth 0.2.5
wheel 0.38.4
zipp 3.15.0
It was super difficut.